글로벌 연구동향
방사선종양학
- 2022년 01월호
[Biomark Med.] Machine-learning algorithms predict breast cancer patient survival from UK Biobank whole-exome sequencing data서울의대 / 장범섭 김인아*
- 출처
- Biomark Med.
- 등재일
- 2021 Nov
- 저널이슈번호
- 15(16):1529-1539. doi: 10.2217/bmm-2021-0280. Epub 2021 Oct 15.
- 내용
Abstract
Aim: We tested whether machine-learning algorithm could find biomarkers predicting overall survival in breast cancer patients using blood-based whole-exome sequencing data. Materials & methods: Whole-exome sequencing data derived from 1181 female breast cancer patients within the UK Biobank was collected. We found feature genes (n = 50) regarding total mutation burden using the long short-term memory model. Then, we developed the XGBoost survival model with selected feature genes. Results: The XGBoost survival model performed acceptably, with a concordance index of 0.75 and a scaled Brier score of 0.146 in terms of overall survival prediction. The high-mutation group exhibited inferior overall survival compared with the low-mutation group in patients ≥56 years (log-rank test, p = 0.042). Conclusion: We showed that machine-learning algorithms can be used to predict overall survival in breast cancer patients from blood-based whole-exome sequencing data.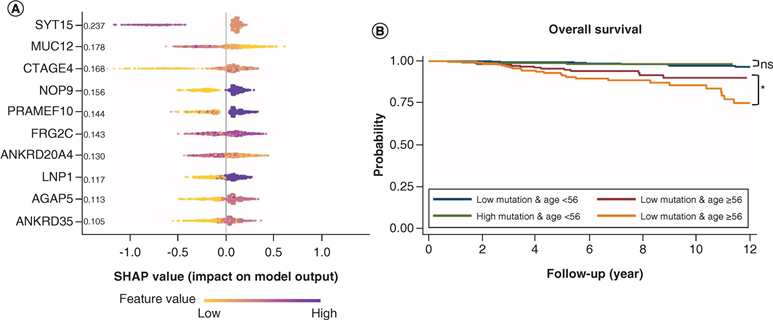
Affiliations
Bum-Sup Jang 1 2 , In Ah Kim 1 2 3
1 Department of Radiation Oncology, Seoul National University Bundang Hospital, Seongnam, 13620, Korea.
2 Department of Radiation Oncology, Seoul National University, College of Medicine, Seoul, Korea.
3 Integrated Major in Innovative Medical Science, Seoul National University Graduate School, Seoul, Korea.
- 키워드
- UK Biobank; breast cancer; machine learning; whole-exome sequencing.
- 연구소개
- UK biobank 내 연구대상자 중 유방암 환자를 대상으로 혈액 샘플에 대한 Whole exome 데이터에 대하여 머신러닝 알고리즘을 적용하여 생존에 대한 예후인자를 밝힌 연구입니다. 혈액 내에 존재하는 COSMIC db기반 variation 혹은 mutation에 대한 정보가 유방암 환자의 생존에 있어 나이와 함께 중요하다는 것이 결론입니다.
- 덧글달기